
Contact
- hnwu@gate.sinica.edu.tw
- 3517, Institute of Molecular Biology, Academia Sinica
Research
The Molecular Mechanism Underlying Dengue Virus Replication and Virion Assembly
Dengue virus (DENV), an enveloped, positive-stranded RNA virus of the Flaviviridae family, is one of the most significant mosquito-borne human viral pathogens. The DENV genome consists of a 5’-capped RNA of ~11 kb that encodes three structural proteins and seven non-structural (NS) proteins. Flavivirus non-structural proteins engage in complicated protein-protein interactions for viral RNA replication machinery assembly. Flavivirus morphogenesis is coupled with viral RNA replication and several flavivirus non-structural proteins have been shown to regulate virion assembly.
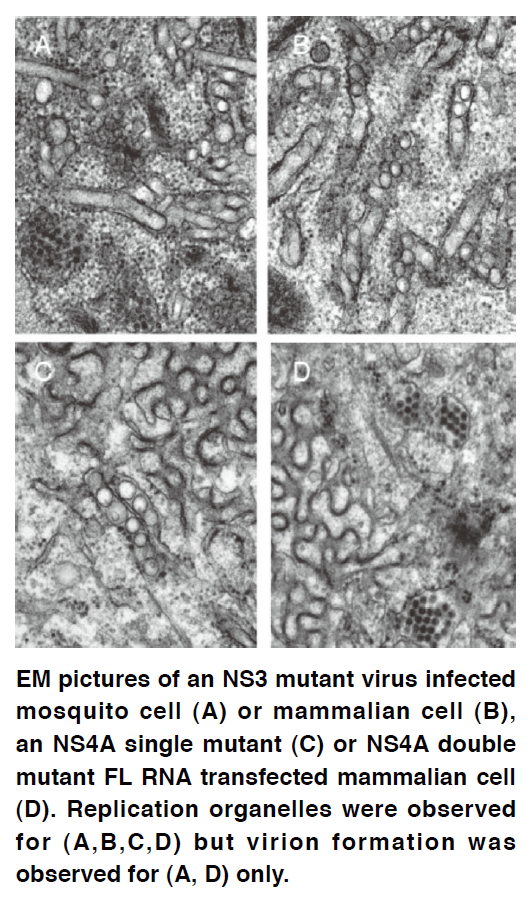
Flaviviral replicons and full-length infectious clones that replicate efficiently in cultured mammalian and mosquito cell lines have been widely used as a tool for reverse and forward genetic analysis of viral gene function. We constructed several DENV replicon systems for basic and application research. A cell line harboring a reporter DENV replicon has been used for anti viral agent screening, which leads to the discovery of several anti DENV compounds by four research groups. A selectable DENV replicon was found to be eligible for performing mutagenesis analysis of viral genes and RNA elements and for screening of suppressor/ adaptive mutation of replication impaired primary mutation. Based on our findings, we built a genetic interaction network of DENV non-structural proteins for replication complex constitution, and established a model for the arrangement of cytosolic NS3 and NS5 proteins, ER lumen-located NS1 protein, and ER membrane-anchored NS4A and NS4B proteins in replication complex. Our study using a DENV infectious clone revealed several DENV non-structural proteins have a role beyond viral RNA replication. We are studying the role of NS4A protein in virion morphogenesis and the possible molecular mechanism, and we are investigating the interplay of NS3 protein with viral structural proteins and non-structural proteins in processes beyond viral RNA replication.
Copyright © 2017 IMBCC. All rights reserved. |