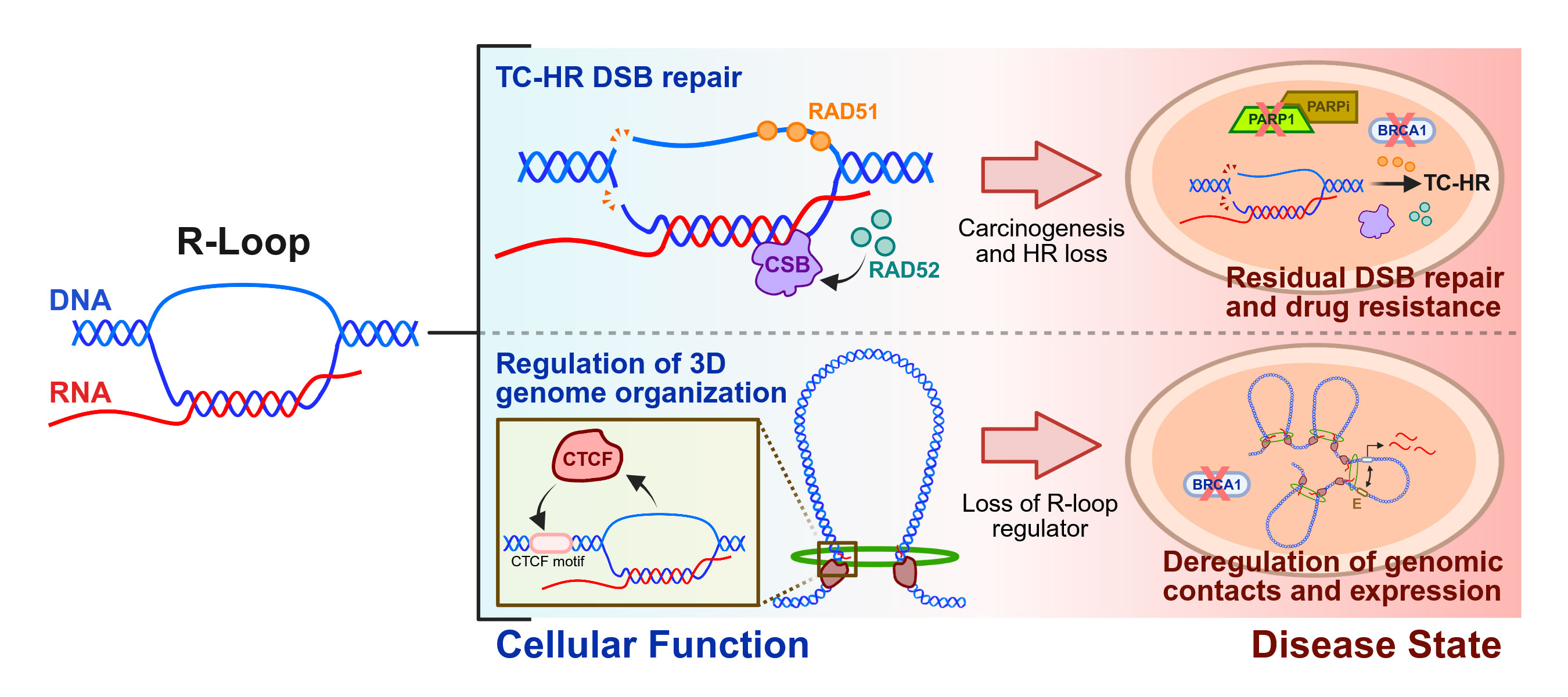
Roles of R-Loop Structures in Cellular Function and Disease Progression
Our group's research focuses on R-loops , dynamic and fascinating three-stranded nucleic acid structures that form within the genomic DNA of all cells. R-loops pose barriers to DNA replication, and their improper accumulation or persistence is thus thought to contribute to disease by causing DNA damage and genome instability. At the same time, however, an abundance of evidence has revealed that these structures play essential roles in normal cellular homeostasis as well. We aim to further uncover these cellular functions and explore how their dysregulation contributes to pathogenesis and disease progression.
One fundamental aspect of our research is identifying cellular interactors of R-loops . We are particularly interested in R-loop "readers": enzymes that bind and/or recognize R-loops in order to connect them to downstream pathways such as gene regulation, chromatin remodeling, or DNA repair. Identifying these readers is crucial to understanding how R-loops are integrated into their cellular roles; elucidating how mutations in readers can impact R-loop mediated function; and developing therapies that target R-loop related vulnerabilities in disease. We utilize innovative proximity labeling strategies, proteomic screens, and cell knockout models to answer these questions.
We are also exploring an emerging and exciting role for R-loops in the regulation of genome organization . R-loops can bolster the binding of CTCF, a master chromatin architectural protein that facilitates 3D organization of the genome at multiple scales. This recent discovery opens up the tantalizing possibility that R-loop accumulation, in addition to consequences on DNA damage and genome stability, may contribute to disease progression by rewiring the 3D genome into a configuration that supports pathogenic gene expression. We are examining this hypothesis by combining cutting-edge genomic sequencing methods to probe how R-loops, CTCF, and chromatin conformation change in tandem upon perturbation in disease models.
Our experimental approaches interface molecular biology with high-throughput genomic sequencing, generating and using cell lines to model R-loop dynamics in disease with a focus on breast and ovarian cancer. Lab members receive extensive depth and breadth of training in the full bench-to-computation pipeline of genomics work, encompassing both wet-lab experimental techniques and dry-lab bioinformatic analysis. Our research will shed light on the fundamentals of R-loop mediated cellular regulation and identify therapeutic targets for diseases and disorders in which R-loops accumulate.
- PDF, 2019-2025, Genome Regulation and Cell Signaling Program, The Wistar Institute
- Ph.D., 2018, Cellular and Molecular Medicine, Johns Hopkins University
- BS, 2008-2012, Human Biology, University of California San Diego
- Bieluszewska A*, Wulfridge P, Fang KC, Hong Y, Sawada T, Erwin J, Song H, Ming GL, Sarma K (2025) Transcriptomic Analysis Uncovers an Unfolded Protein Response in ADNP Syndrome. Molecular and Cellular Biology 45(4):143-153
- Grothusen GP*, Chang R, Cao Z, Zhou N, Mittal M, Datta A, Wulfridge P, Beer T, Wang B, Zhen N, Tang HY, Sarma K, Greenberg RA, Shi J, Busino L (2024) DCAF15 control of cohesin dynamics sustains acute myeloid leukemia. Nature Communications 15(1):5604
- Wulfridge P*, Davidovich A*, Salvador AC*, Manno GC, Tryggvadottir R, Idrizi A, Huda MN, Bennett BJ, Adams LG, Hansen KD, Threadgill DW, Feinberg AP (2023) Precision pharmacological reversal of strain-specific diet-induced metabolic syndrome in mice informed by epigenetic and transcriptional regulation. PLoS Genetics 19(10):e1010997
- Wulfridge P*, Yan Q, Rell N, Doherty J, Jacobson S, Offley S, Deliard S, Feng K, Phillips-Cremins JE, Gardini A, Sarma K (2023) G-quadruplexes associated with R-loops promote CTCF binding. Molecular Cell 83(17):3064-3079.e5
- Bieluszewska A, Wulfridge P, Doherty J, Ren W, Sarma K (2022) ATRX histone binding and helicase activities have distinct roles in neuronal differentiation. Nucleic Acids Research 50(16):9162-9174
- Yan Q*, Wulfridge P*, Doherty J, Fernandez-Luna JL, Real PJ, Tang HY, Sarma K (2022) Proximity labeling identifies a repertoire of site-specific R-loop modulators. Nature Communications 13(1):53
- Wulfridge P*, Sarma K (2021) A nuclease- and bisulfite-based strategy captures strand-specific R-loops genome-wide. eLife 10:e65146
- Wulfridge P*, Langmead B, Feinberg AP, Hansen KD (2019) Analyzing whole genome bisulfite sequencing data from highly divergent genotypes. Nucleic Acids Research 47(19):e117
- Barrington WT*, Wulfridge P, Wells AE, Rojas CM., Howe SYF, Perry A, Hua K, Pellizzon MA, Hansen KD, Voy BH, Bennett BJ, Pomp D, Feinberg AP, Threadgill DW (2018) Improving Metabolic Health Through Precision Dietetics in Mice. Genetics 208(1):399-417