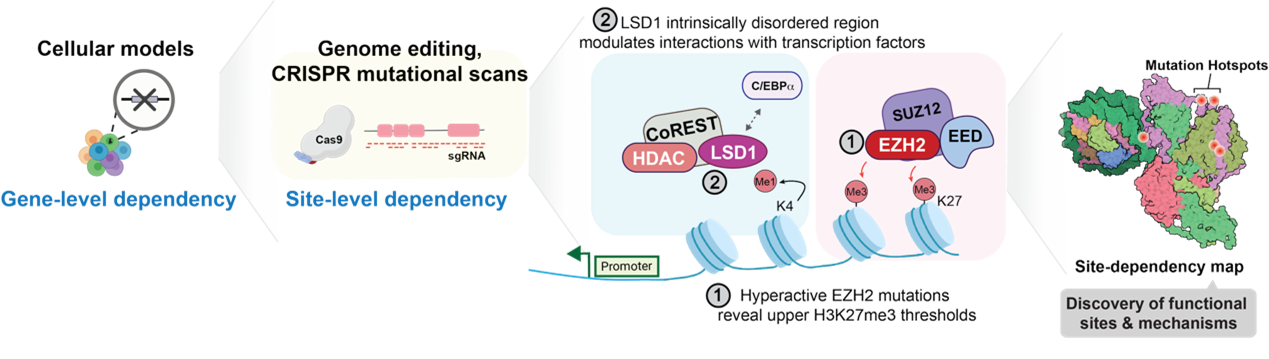
透過功能性基因體學探究基因調控機制
染色質調控因子(Chromatin Regulators),包括組蛋白、修飾酶、轉錄因子以及染色質重塑因子,構成了控制基因體存取的核心分子機制。值得注意的是,編碼這些因子的基因是癌症中最常發生突變的基因類別之一。我們致力於研究染色質相關因子如何調控基因表現,以及其功能失調如何促成癌症的發生。透過功能性基因體學的方法,我們希望揭示染色質調控異常如何促成疾病的發生。
我們致力於理解與疾病相關的染色質調控因子突變如何擾亂基因調控。透過結合大規模基因功能分析與深入的分子機制研究,我們系統性地解析基因調控異常的來源,並連結DNA 序列、分子機制與細胞表型,目標在於發掘具治療潛力的癌症脆弱點。
我們透過以下研究架構來實現上述目標:
- 探索 (Discovery): 鑑定在染色質調控因子中具有功能性與細胞層級影響的突變
- 機制 (Mechanism): 解析關鍵突變如何在分子層級改變染色質功能
- 臨床轉譯 (Translation): 將分子機制連結至細胞表型,並鑑定具治療潛力的標的與策略
我們關注在癌症中常見的離胺酸甲基轉移酶複合體與組蛋白突變,並探討這些變異如何促成疾病的發生。
- PDF, 2019-2025, Department of Chemistry and Chemical Biology, Harvard University
- PDF, 2017-2019, Department of Molecular Biophysics and Biochemistry, Yale University
- Ph.D., 2011-2016, NUS Graduate School – Integrative Sciences and Engineering Program, National University of Singapore (NUS)
- B.S., 2007-2010, School of Biological Sciences, Nanyang Technological University, Singapore
- 2025, Forbeck Foundation Scholar Award
- 2023, Abstract Achievement Award, American Society of Hematology (ASH).
- 2023, Eliana Hechter Memorial Travel Prize, Broad Institute.
- 2022, Charles A. King Trust Postdoctoral Research Fellowship, Health Resources in Action.
- Kwok HS*, Morriss JW*, Hu CXY*, Iram I, Myung Y, Siegenfeld AP, Waterbury AL, Yeo MJR, Lee C, Camp S, Iqbal S, Liau BB. (2025) Base editing charts a sequence-function atlas of Polycomb Repressive Complex 2 at amino acid resolution. Submitted.
- Kwok HS*, Freedy AM*, Siegenfeld AP, Morriss JW, Waterbury AL, Kissler SM, Liau BB. (2023) Drug addiction mutations unveil a methylation ceiling in EZH2-mutant lymphoma. Nature Chemical Biology. 19, 1105-1115. doi: 10.1038/s41589-023-01299-1.
- Waterbury AL*, Kwok HS*, Lee C*, Narducci D, Freedy AM, Su C, Raval S, Reiter A, Hawkins W, Lee K, Li J, Hoenig SM, Vinyard ME, Cole PA, Hansen AS, Carr SA, Papanastasiou M, Liau BB. (2024) An autoinhibitory switch of the LSD1 disordered region controls enhancer silencing. Molecular Cell, 84, 2238-2254. doi: 10.1016/j.molcel.2024.05.017
- Yeo MJR*, Zhang O*, Xie X*, Nam E*, Payne NC, Gosavi PM, Kwok HS, Iram I, Lee C, Li J, Chen N, Nguyen K, Jiang H, Wang ZA, Lee K, Mao H, Harry SA, Barakat I, Takahashi M, Waterbury AL, Barone M, Mattevi A, Carr SA, Udeshi ND, Bar-Peled L, Cole PA, Mazitschek R, Liau BB.#, Zheng N.# (2025) UM171 glues asymmetric CRL3–HDAC1/2 assembly to degrade CoREST corepressors. Nature, 639, 232–240. doi.org/10.1038/s41586-024-08532-4 (#Co-corresponding authors)
- Xie X*, Zhang O*, Yeo MJR*, Lee C*, Tao R, Harry SA, Payne NC, Nam E, Paul L, Li Y, Kwok HS, Jiang H, Mao H, Hadley JL, Lin H, Batts M, Gosavi PM, D’Angiolella V, Cole PA, Mazitschek R, Northcott PA, Zheng N#, Liau BB#. (2025) Converging mechanism of UM171 and KBTBD4 neomorphic cancer mutations. Nature, 639, 241–249. doi.org/10.1038/s41586-024-08533-3 (#Co-corresponding authors)
- Kwok HS#, Vargas-Rodriguez O, Melnikov SV, Söll D. (2019) Engineered aminoacyl-tRNA synthetases with improved selectivity toward noncanonical amino acids. ACS Chemical Biology, 14(4), 603–612. doi: 10.1021/acschembio.9b00088 (# Sole corresponding author)
- Numata A*, Kwok HS*, Zhou QL*, Li J, Tirado-Magallanes R, Angarica VE, Hannah R, Park J, Wang CQ, Krishnan, V, Rajagopalan D, Zhang Y, Zhou S, Welner RS, Osato M, Jha S, Bohlander SK, Göttgens B, Yang H, Benoukraf T, Lough JW, Bararia D, Tenen DG. (2020) Lysine acetyltransferase Tip60 is required for hematopoietic stem cell maintenance. Blood, 136(15), 1735–1747. doi: 10.1182/blood.2019001279
- Numata A*, Kwok HS*, Kawasaki A, Li J, Zhou QL, Kerry J, Benoukraf T, Bararia D, Li F, Ballabio E, Tapia M, Deshpande AJ, Welner RS, Delwel R, Yang H, Milne TA, Taneja R, Tenen DG. (2018) The basic helix-loop-helix transcription factor SHARP1 is an oncogenic driver in MLL-AF6 acute myelogenous leukemia. Nature Communications, 9(1), 1622. doi: 10.1038/s41467-018-03854-0
- Bararia D*, Kwok HS*, Welner RS, Numata A, Sárosi MB, Yang H, Wee S, Tschuri S, Ray D, Weigert O, Levantini E, Ebralidze AK, Gunaratne J, Tenen DG. (2016) Acetylation of C/EBPα inhibits its granulopoietic function. Nature Communications, 7(1), 10968. doi: 10.1038/ncomms1096
* denotes co-first authors, # denotes corresponding authors