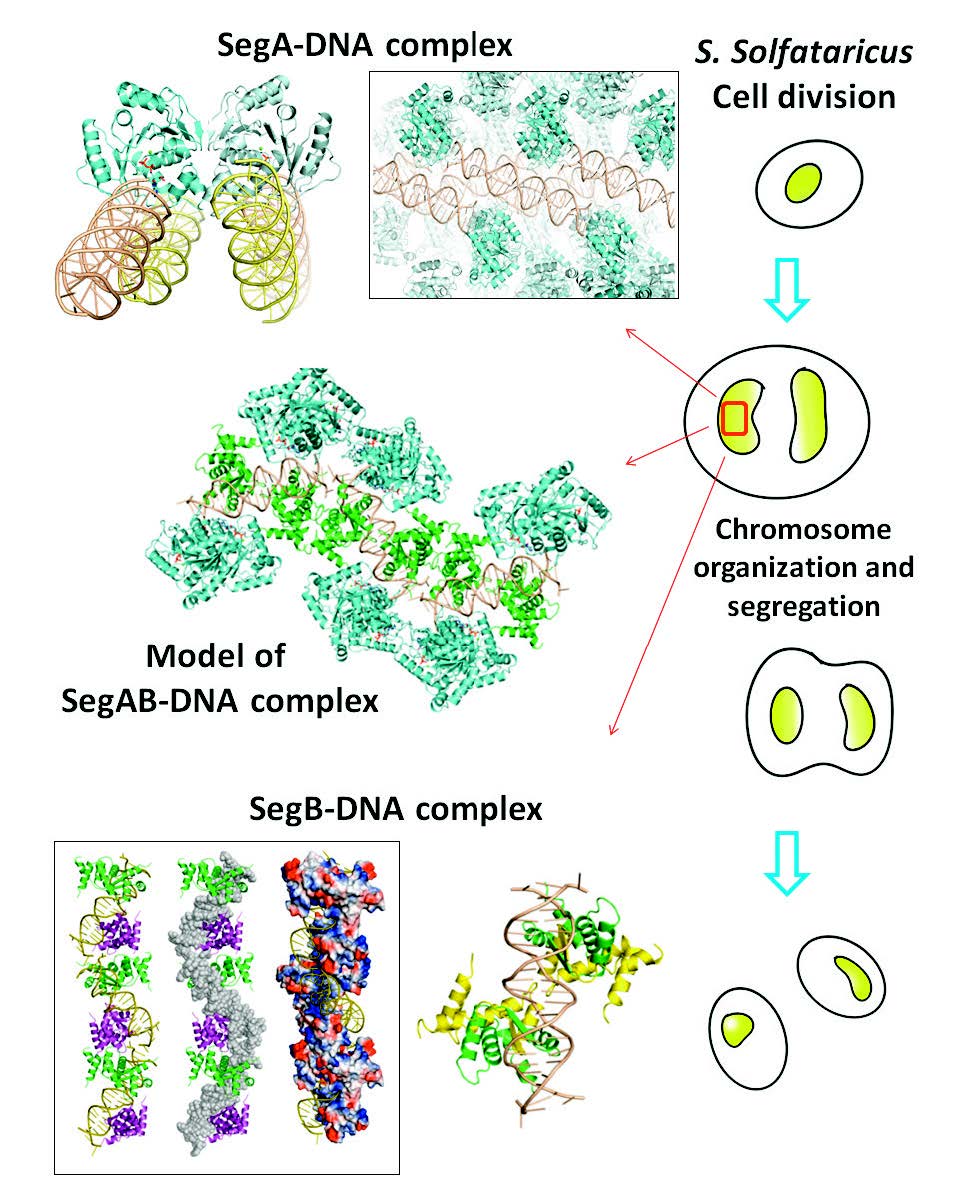
The Structure and Mechanism of Chromosome Segregation
The newly replicated DNA is required further segregation or partition to ensure that the genetic material is maintained. Many protein complexes play a vital role in this process. In eukaryotes, the replicated chromosomes are segregated through mitoic spindle, but the plasmid ParABS cassette is the most used protein complex for bacterial chromosome segregation. However, the genomes segregation is still unclear in archaea, the thirddomain of life. Recently, SegA and SegB, the components participate in the archaeal chromosome segregation have been identified and have fundamental realization. However, the detailed mechanism and the process of archaeal chromosome separation are still unknown. In this proposal, we will combine biological functional and biophysical techniques to investigate the structure and mechanism of proteins involved in this segregation system. Information from this study, we hope to fill up the missing piece of chromosome segregation in the third kingdom of life and to illustrate the mechanism of archaeal chromosome segregation.
Structural and Functional Study of the Protein Involved in DNA Replication Restart
Precise and reliable transmission of genetic infor-mation is critical to the cell proliferation in all organ- isms. However, when cells are stressed by environmental damaging treatments, DNA replication forks often encounter template DNA lesions, which can stall the progression of the replication forks. For surviving, cells must be able to reactivate replisomes that are disrupted in these reasons. In bacteria, this process is termed ‘‘DNA replication restart’’ and it is governed by a group of proteins called primosome proteins. In this proposal, we will study the relationship between structure and function of these components by X-ray crystallography, molecular biology and biochemical approaches.
- PDF, 1993-1995, IMB, Academia Sinica
- Ph.D., 1993, Dept.Crystallography, Univ. of Pittsburgh, USA
- MS, 1984, Dept.Chemistry, Natl. Taiwan Univ.
- BS, 1982, Dept. Chemistry, Chung-Yuan Christian Univ.
- 2000, Academia Sinica Early-Career Investigator Research Achievement Award
- 2002, Research Excellence Award, National Science and Technology Council
- 2007-2011, Academia Sinica Investigator Award
- 2021, Academia Sinica Presidential Scholars Program
- Sun, Y.-J., Forouhar, F., Li, H.-M., Tu, S.-L., Kao, S., Shr, H.-L., Chou, C.-C., Hsiao, C.-D. (2002) Crystal structure of pea Toc34 - a novel GTPase of the chloroplast protein translocon. Nature Structure Biology 9: 95-100.
- Yeh, Y.H., Lin, T., Lin, C.Y. Hsiao, C.D. (2014) Crystal structure and functional characterizations of Ybr137w from Saccharomyces cerevisiae. Mol. Cell Biol. 34: 4500-4512.
- Lou, Y.C., Weng, T.H., Li, Y.C., Kao, Y.F., Chou, S.H., *Hsiao, C.D., *Chen, C. (2015) Structure and dynamics of the polymyxin-resistance-associated response regulator PmrA in complex with the promoter DNA. Nature Communications 6: 8838 doi:10.1038/ ncomms9838.
- Li, Y.C., Naveen, V., Lin, M.G.,Hsiao, C.D. (2017) Structural analyses of the bacterial primosomal protein DnaB reveal that it is a tetramer and forms a complex with a primosomal re-initiation protein. J. Biol. Chem. 292(38): 15744-15757.
- Chu, C.H., Yen, C.Y., Chen, B.W., Lin, M.G., Wang, L.H., Tang, K.Z., *Hsiao, C.D., *Sun, Y.J. (2019) Crystal structures of HpSoj–DNA complexes and the nucleoidadaptor complex formation in chromosome segregation. Nucleic Acids Res. 47(4): 2113-2129.
- Tsai, J.Y., Chu, C.H., Lin, M.G., Chou, Y.H., Hong, R.Y., Yen, C.Y., *Hsiao, C.D., *Sun, Y.J. (2020) Structure of the sodium-dependent phosphate transporter reveals insights into human solute carrier SLC20. Science Advances 7: 6(32): eabb4024.