Meiosis
Meiosis, a cell division process unique to the sexual reproductive cells of eukaryotes, functions to halve the number of chromosomes, producing haploid gametes (sperm and eggs) or fungal sexual spores. Meiosis is important for two reasons. First, it creates “genomic diversity” in a species through three primary mechanisms: DNA repair and homologous recombination (HR) between homologous non-sister chromosomes in prophase I; independent assortment of homologous chromosomes in metaphase I; and random fertilization. Second, meiosis is closely linked to speciation. Postzygotic hybrid infertility is a mechanism of reproductive isolation whereby the hybrid offspring of two closely related species survive to adulthood but cannot complete meiosis and generate fertile gametes or viable spores, thus maintaining distinct species boundaries. Our laboratory uses the budding yeast Saccharomyces cerevisiae and an industrial workhorse filamentous fungus Trichoderma reesei as model organisms to investigate several important meiotic processes, including DNA metabolism (i.e., replication, repair, and recombination), chromosome dynamics, genetic and epigenetic regulation, as well as the evolutionary diversity of meiosis. We are also interested in understanding the molecular mechanisms and functions of “chromosome individualization”, representing a critical process during cell division (mitosis/meiosis) whereby condensed chromosomes separate from one another, preventing tangling and ensuring accurate segregation into daughter cells.
Homologous recombination counteracts mismatch repair to promote fertility and genetic diversity
The proteins involved in DNA repair and homologous recombination (HR), as well as the ZMM proteins that make up the synaptonemal complex (SC), can inhibit the activity of mismatch repair (MMR). The SC is a meiosis-specific, zipper-like tripartite structure that connects homologous chromosomes during meiotic prophase. We examined whole-genome research data and other relevant literature to explain the fallacies in previous theories regarding "MMR causing hybridization infertility and reproductive isolation in closely related species" present in biology and genetics textbooks over the past 30 years. Next, using "reverse genetics", we identified 12 HR mutant genes that specifically affect the fertility of the hybrid zygotes generated by mating two intraspecies S. cerevisiae species, SK1 and S288c. The same mutant genes do not or only slightly affect the meiosis of SK1/SK1 purebred zygotes, demonstrating that multiple HR and SC proteins can inhibit MMR activity and tolerate mismatched DNA bases during meiotic recombination. Our new discovery is important for two reasons. First, it challenges the prevailing theory that “HR is the mechanism for repairing damaged DNA with the highest fidelity”. Second, though the SC has been known for nearly 70 years, its physiological function had remained unknown. We demonstrated for the first time that the SC is responsible for suppressing MMR during hybrid meiosis, thus promoting recombination and exchange between homologous chromosomes with different DNA sequences.
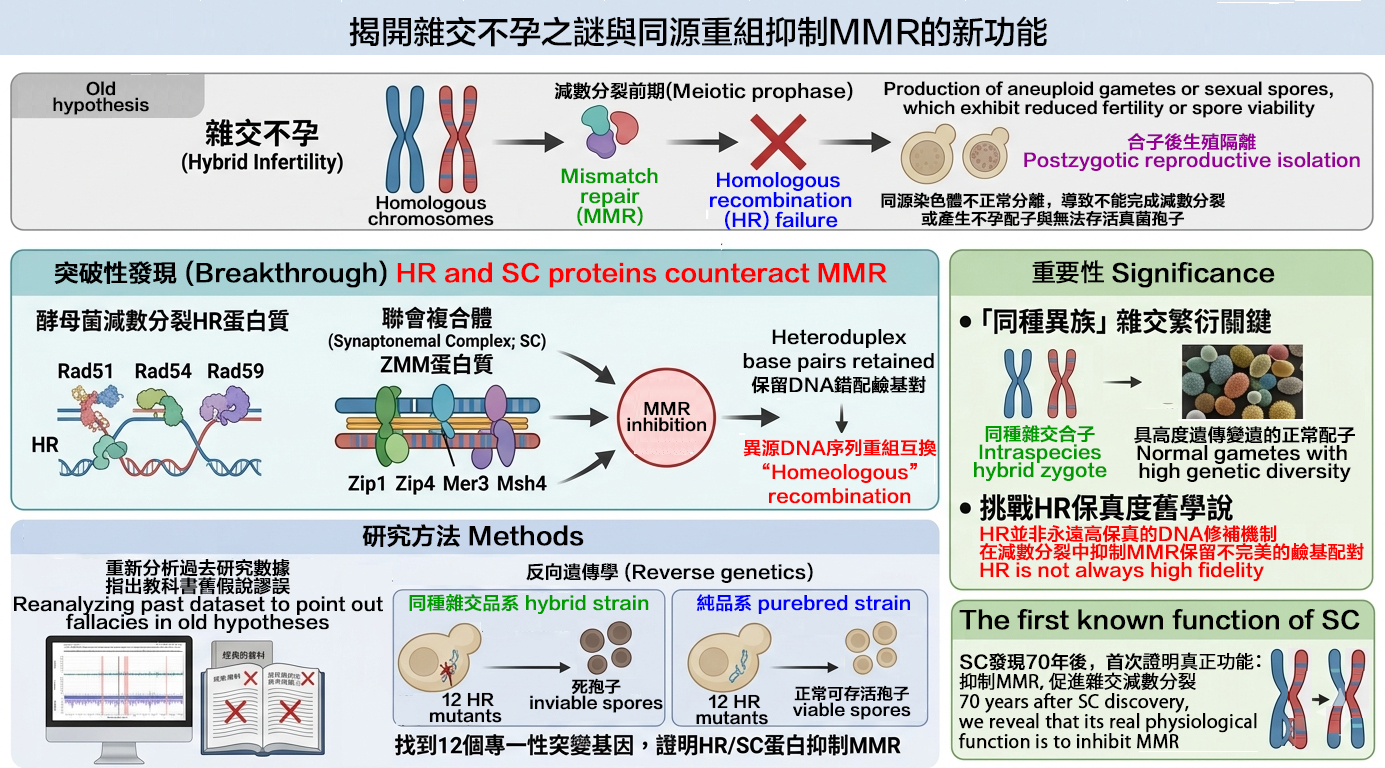
- *Ting-Fang Wang, and Ji-Long Liao (2026) Homologous recombination counteracts mismatch repair to promote fertility and genetic diversity. Trends in Genetics 42, 7-10. PMID: 41241584. (*corresponding author).
- #Ya-Ling Hung, #Chin-Ning Chuang, Hong-Xian Kim, Hou-Cheng Liu, Jhong-Syun Yao, Lavernchy Javanska, and *Ting-Fang Wang (2025) Rad51, Rad54, and ZMM proteins antagonize the mismatch repair system to promote fertility of budding yeast intraspecies hybrid zygotes. Nucleic Acids Research 53, gkaf847. PMID: 40902007 (#contributed equally).
- Ta-Ting Woo, Chi-Ning Chuang, Mika Higashide, Akira Shinohara, and *Ting-Fang Wang (2020) Dual roles of yeast Rad51 N-terminal domain in repairing DNA double-strand breaks. Nucleic Acids Research 48, 8474-8489 PMID:32652040.
Genetic and epigenetic regulation of Trichoderma reesei meiosis
In many sexually reproducing eukaryotes (such as budding yeast S. cerevisiae, fission yeast S. pombe, higher plants, and mammals), meiotic cells utilize the Spo11 protein to generate DNA double-strand breaks (DSBs) and two RecA-like recombinases (Rad51 and meiosis-specific Dmc1) to repair DSBs, thereby promoting genetic exchange between homologous non-sister chromosomes and generating new genetic diversity in the genomes of gametes (sperm and eggs) or fungal sexual spores. Mammalian and yeast Dmc1 can stabilize recombination intermediates containing mismatches better than Rad51. We discovered that T. reesei possesses three unique meiotic features. First, the T. reesei spo11 gene is dispensable for meiosis and the production of fertile (or viable) sexual spores. Second, T. reesei only has Rad51 but not Dmc1. Rad51 is indispensable for interhomolog recombination during T. reesei meiosis. Like S. cerevisiae Dmc1, T. reesei Rad51 possesses better mismatch tolerance than S. cerevisiae Rad51 during meiosis. The ancestral T. reesei Rad51 evolved to acquire Dmc1-like properties by creating multiple structural variations, including via amino acid residues in the L1 and L2 DNA-binding loops. Third, T. reesei DNA methyltransferases (DMTs: RID1, DIM2, and DIMX) exert multiple functions in meiosis: (1) RID1 and DIM2 are responsible for initiating meiosis or meiotic entry; (2) RID1 promotes DNA repair during meiotic recombination, thereby promoting new genetic variations; and (3) through a special epigenetic mechanism, RID1, DIM2 and DIMX methylate a large amount of cytosine on the parental DNA, inducing point mutations to thymine, rapidly increasing the genomic diversity of meiotic products (sexual spores). Excessive genetic variation has double-edged effects. Although it can enhance the environmental adaptability of offspring, it often leads to genomic instability, preventing sexual spores from germinating into mycelia or inducing newly germinated mycelia to fail to undergo normal vegetative growth. Our results explain why many Trichoderma wild-isolates are female- or male-infertile. They can only reproduce asexually through vegetative growth of mycelium, which actually helps maintain genomic stability and perpetuate their existing survival advantages.
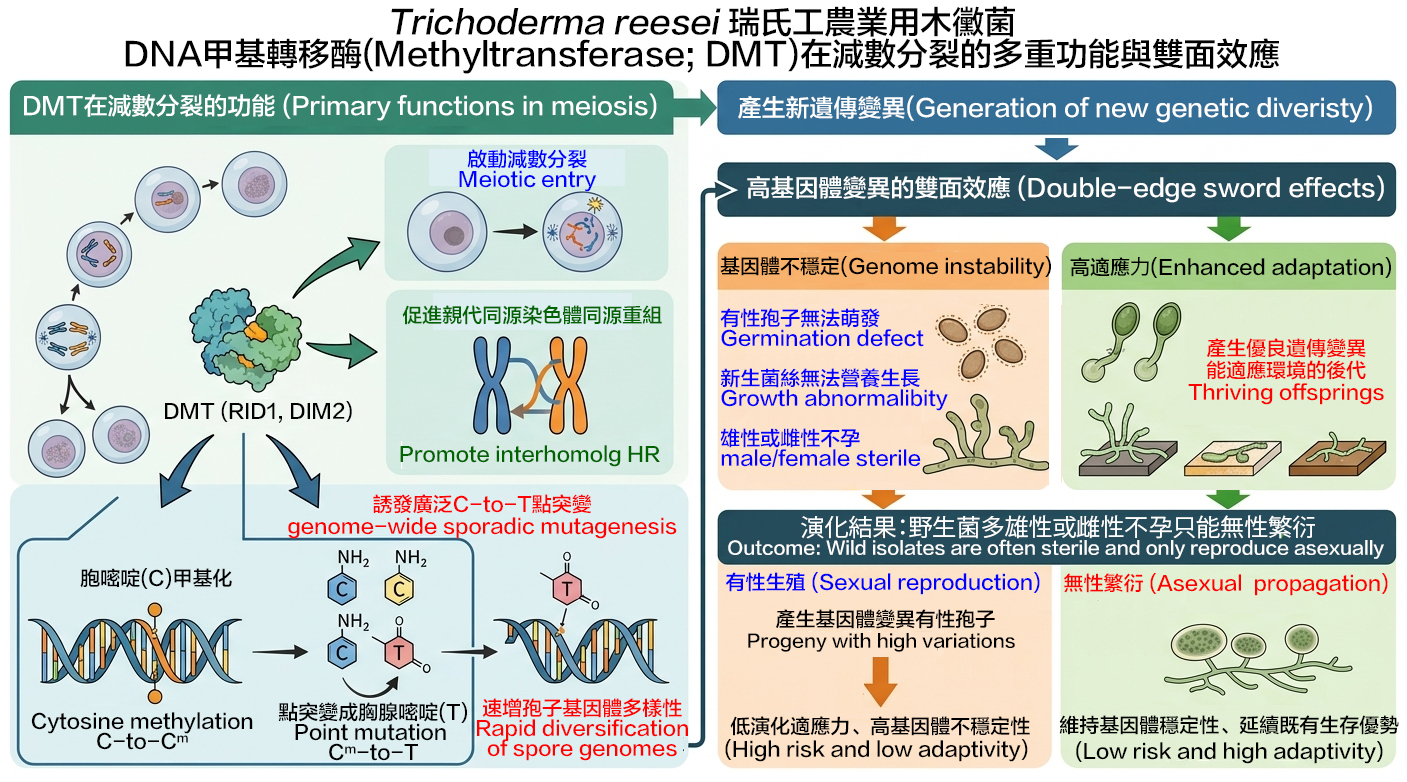
- #Lavernchy Jovanska, #I-Chen Lin, Jhong-Syuan Yao, Chia-Ling Chen, Hou-Cheng Liu, Wan-Chen Li, Yu-Chien Chuang, Albert Chen-Hsin Yu, Hisn-Nan Lin, Wen-Li Peng, Chang-I Yu, Ching-Yuan Su, Yi-Ping Chen, Ruey-Shyand Chen, Yi-Ping Hsueh, Hanna S Yuan, Ljudmilla Timofejeve, and *Ting-Fang Wang (2024) DNA cytosine methyltransferase differentially regulate genome-wide hypermutation and interhomolog recombination in Trichoderma reesei. Nucleic Acids Research 52, 9551-9537. PMID: 39021337.
- #Wan-Chen Li, #Chia-Yi Lee, #Wei-Hsuan Lan, #Tai-Ting Woo, Hou-Cheng Liu, Hsin-Yi Yen, Hao-Yen Chang, Yu-Chien Chuang, Chiung-Ya Chen, Chi-Ning Chuang, Chia-Ling Chen, Yi-Ping Hsueh, *Hung-Wen Li, *Peter Chi, and *Ting-Fang Wang (2021) Trichoderma reesei Rad51 tolerates mismatches in hybrid meiosis with diverse genome sequences. Proc. Natl. Acad. Sci. USA 118, e2007192118. PMID: 33593897.
- Hou-Cheng Liu, Wan-Chen Li, and *Ting-Fang Wang (2021) TSETA: a third-generation-sequencing based program for single-nucleotide-resolution mapping of meiotic recombination products and repeat-induced point mutations. Methods in Molecular Biology 2234, 331-361. PMID: 3316579.
- Wan-Chen Li, and *Ting-Fang Wang (2021) PacBio long-read sequencing, assembly, and Funannotate reannotation of the complete genome of Trichoderma reesei QM6a. Methods in Molecular Biology 2234, 311-329. PMID: 33165795.
The nuclear nonhistone protein Ki-67 regulates perichromosomal lipids in mammalian cells
Nuclear lipids play roles in regulatory processes such as signaling, transcriptional regulation, and DNA repair. We report that nuclear lipids may contribute to Ki-67-regulated chromosome integrity during mitosis. In COS-7 cells, nuclear lipids are enriched at the perichromosomal layer and excluded from intrachromosomal regions during early mitosis, but are then detected in intrachromosomal regions during late mitosis. The nuclear non-histone protein Ki-67 acts as a surfactant to form a repulsive molecular brush around fully condensed sister chromatids in early mitosis, preventing the diffusion or penetration of nuclear lipids into intrachromosomal regions. Ki-67 is phosphorylated during mitosis by cyclin-dependent kinase 1 (CDK1), the best-known master regulator of the cell cycle. Both Ki-67 knockdown and reduced Ki-67 phosphorylation by CDK1 inhibitors allow nuclear lipids to penetrate chromosomal regions. Thus, both Ki-67 protein level and phosphorylation status during mitosis appear to influence the perichromosomal distribution of nuclear lipids. Ki-67 knockdown and CDK1 inhibition also lead to uneven chromosome disjunction between daughter cells, highlighting the critical role of this regulatory mechanism in ensuring accurate chromosome segregation. Given that Ki-67 has been proposed to promote chromosome individualization and establish chromosome-cytoplasmic compartmentalization during open mitosis in vertebrates, our results reveal that nuclear lipid enrichment at the perichromosomal layer enhances Ki-67’s ability to form a protective perichromosomal barrier (chromosome envelope; CE), which is critical for correct chromosome segregation and maintenance of genome integrity during mitosis.
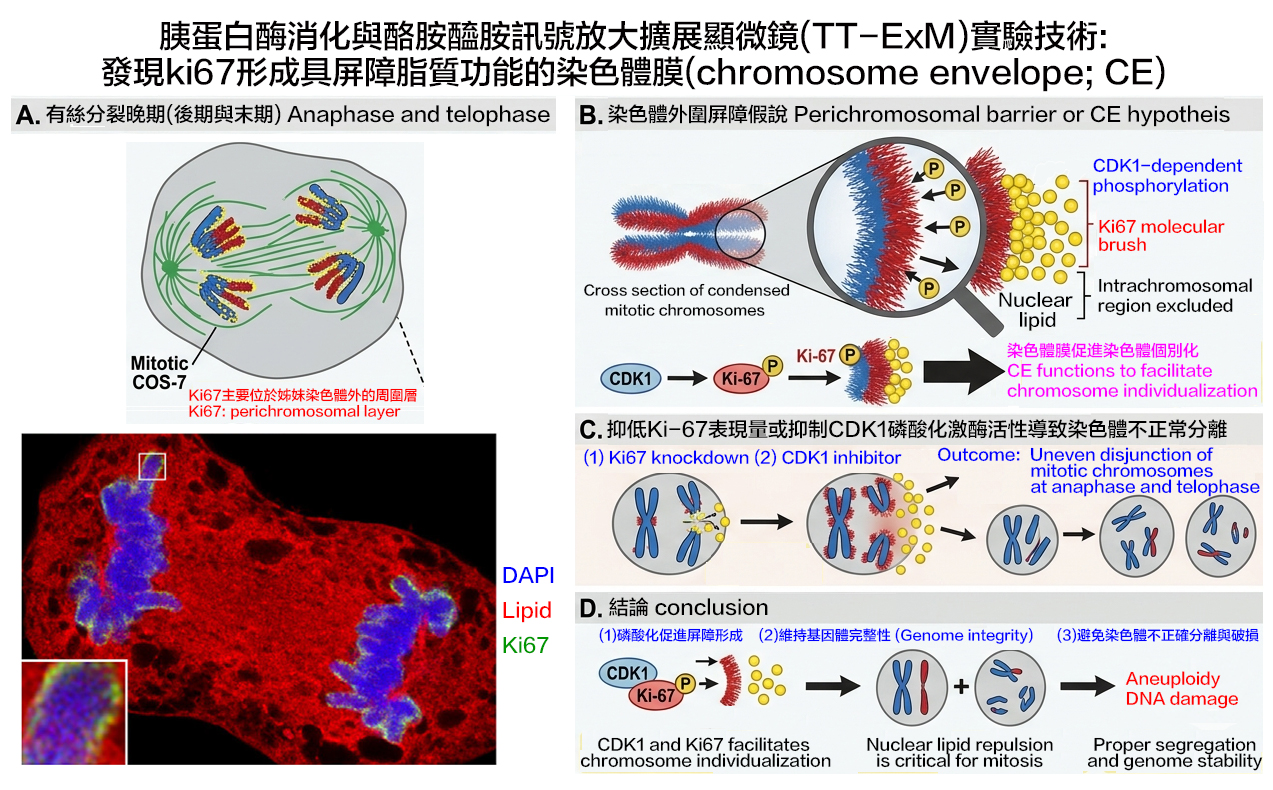
- Hsiao-Tang Hu, Ueh-Ting Tim Wang, Bi-Chang Chen, *Yi-Ping Hsueh, and *Ting-Fang Wang (2025) Ki-67 and CDK1 control the dynamic association of nuclear lipids with mitotic chromosomes. Journal of Lipid Research 66, 100731. (This paper has been downloaded 1327 times since January, 2025, and was supported in part by an intramural grant (SPP) from IMB, Academia Sinica).
- PDF, 1999-2000, Dept. Mol. & Cell Biol., Harvard Univ. USA
- Ph.D., 1998, Dept. Mol. & Cell Biol., Harvard Univ. USA
- MS, 1990, Inst. Life Science., Natl. Tsing Hua Univ.
- BS, 1988, Dept. Chemistry, Natl. Taiwan Univ.
- Ting-Fang Wang# and Ji-Long Liao. (2025) Homologous recombination counteracts mismatch repair to promote fertility and genetic diversity. Trends in Genetics (PMID: 41241584; DOI: 10.1016/j.tig.2025.10.009)
- Ya-Ling Hung, Chi-Ning Chuang, Hong-Xiang Kim, Hou-Cheng Liu, Jhong-Syuan Yao, Lavernchy Jovanska, Yi-Ping Hsueh, Ruey-Shyang Chen, and Ting-Fang Wang# (2025) Rad51, Rad54, and ZMM proteins antagonize the mismatch repair system to promote fertility of budding yeast intraspecies hybrid zygotes. Nucleic Acids Research (PMID: 40902007; DOI: 10.1093/nar/gkaf847)
- Hsiao-Tang Hu, Ueh-Ting Tim Wang, Bi-Chang Chen, Yi-Ping Hsueh#, Ting-Fang Wang# (2025) Ki-67 and CDK1 control the dynamic association of nuclear lipids with mitotic chromosomes. Journal of Lipid Research ( PMID: 39706365; DOI: 10.1016/j.jlr.2024.100731)
- Lavernchy Jovanska, I-Chen Lin, Jhong-Syuan Yao, Chia-Ling Chen, Hou-Cheng Liu, Wan-Chen Li, Yu-Chien Chuang, Chi-Ning Chuang, Albert Chen-Hsin Yu, Hsin-Nan Lin, Wen-Li Pong, Chang-I Yu, Ching-Yuan Su, Yi-Ping Chen, Ruey-Shyang Chen, Yi-Ping Hsueh, Hanna S Yuan, Ljudmilla Timofejeva, Ting-Fang Wang# (2024) DNA cytosine methyltransferases differentially regulate genome-wide hypermutation and interhomolog recombination in Trichoderma reesei meiosis. Nucleic Acids Research (PMID: 39021337; DOI: 10.1093/nar/gkae611)
- Chi-Ning Chuang, Hou-Cheng Liu, Tai-Ting Woo, Ju-Lan Chao, Chiung-Ya Chen, Hisao-Tang Hu, Yi-Ping Hsueh, Ting-Fang Wang# (2024) Noncanonical usage of stop codons in ciliates expands proteins with structurally flexible Q-rich motifs. eLife (PMID: 38393970; DOI: 10.7554/eLife.91405)
- Chia-Lin Chen, Wan-Chen Li, Yu-Chien Chuang, Hou-Cheng Liu, Chien-Hao Huang, Ko-Yun Lo, Chung-Yu Chen, Fang-Mo Chang, Guo-An Chang, Yu-Ling Lin, Wen-denr Yang, Ching-Hua Su, Tsung-Ming Yeh, and Ting-Fang Wang# (2022) Sexual crossing, chromosome-level genome sequences, and comparative genomic analyses for the medicinal mushroom Taiwanofungus camphoratus (Syn. Antrodia cinnamomea; Antrodia camphorata). Microbiology spectrum (PMID: 35196809; DOI: 10.1128/spectrum.02032-21)
- Wan-Chen Li, Ting-Chan Lin, Chia-Ling Chen, Hou-Cheng Liu, Hisn-Nan Lin, Ju-Lan Chao, Cheng-Hsilin Hsieh, Hui-Fang Ni, Ruey-Shyang Chen# and Ting-Fang Wang# (2021) Complete genome sequences and genome-wide characterization of Trichoderma biocontrol agents provide new insights into their evolution and variation in genome organization, sexual development, and fungal-plant interactions. Microbiology spectrum (PMID: 34908505; DOI: 10.1128/Spectrum.00663-21)
- Wan-Chen Li, Chia-Yi Lee, Wei-Hsuan Lan, Tai-Ting Woo, Hou-Cheng Liu, Hsin-Yi Yeh, Hao-Yan Chang, Yu-Chien Chuang, Chi-Ning Chuang, Chia-Lin Chen, Yi-Ping Hsueh, Hung-Wen Li#, Peter Chi#, and Ting-Fang Wang# (2021) Trichoderma reesei Rad51 tolerates mismatches in hybrid meiosis with diverse genome sequences. Proceedings of the National Academy of Sciences (PMID: 33593897; DOI: 10.1073/pnas.2007192118)
- Wan-Chen Li and Ting-Fang Wang# (2021) PacBio long-read sequencing, assembly, and Funannotate reannotation of the complete genome of Trichoderma reesei QM6a. Methods in Molecular Biology (PMID: 33165795; DOI: 10.1007/978-1-0716-1048-0_21)
- Hou-Cheng Liu, Wan-Chen Li, and Ting-Fang Wang# (2021) TSETA: A third-generation sequencing-based computational tool for mapping and visualization of SNPs, meiotic recombination products, and RIP Mutations. Methods in Molecular Biology (PMID: 33165796; DOI: 10.1007/978-1-0716-1048-0_2)
- Tai-Ting Woo, Chi-Ning Chuang, Mika Higashide, Akira Shinohara, and Ting-Fang Wang# (2020) Dual roles of yeast Rad51 N-terminal domain in repairing DNA double-strand breaks. Nucleic Acids Research (PMID: 32652040; DOI: 10.1093/nar/gkaa587)
- Wan-Chen Li, Hou-Cheng Liu, Ying-Jyun Lin, Shu-Yun Tung, Ting-Fang Wang# (2020) Third-generation sequencing-based mapping and visualization of single nucleotide polymorphism, meiotic recombination, illegitimate mutation and repeat-induced point mutation. NAR Genomics and Bioinformatics (PMID: 33575607; DOI: 10.1093/nargab/lqaa056)
- Wang, T.-F., Chin-Ning Chuang and Tai-Ting Woo. “Method and vector for enhancing protein expression”. US provisional application (63/036,704), Patent Cooperation Treaty application (PCT/US2021/036247) and Taiwan application (110120541).
- Wang, T.-F., “Method for producing segmental aneuploidy (SAN) strains of Trichoderma reesei via sexual crossing and SAN strains produced therefrom”. Patent Cooperation Treaty (PCT/US14/58654) and US (9598738B2), Taiwan (I570236) and People’s Republic of China (IA001/ACA081CN).
- Wang, T. F. and Yang, S.M. “Methods of producing virus-like particles of piconavirus using a small-ubiquitin-related fusion protein expression system”. US (8663951B2) and Taiwan (I415623).
- Wang, T.-F. “SUMO fusion protein expression system for producing native proteins”. USA 8034910BZ.
Principal Investigator
Ting-Fang Wang
PhD. Student
Ya-Ling Hung
Postdoctoral Fellow
Lavernchy Jovanska . Ji-Long Liu
Research Assistant
Chi-Ning Chuang . Su-Jie Chen . Hong-Xiang Kim . Hou-Cheng Liu . Nien-Chi Wu
