Splicing regulation and intron evolution in the short-intron ciliate model of endosymbiosis Paramecium bursaria
Dr. Lin, Chien-Ling and Leu, Jun-Yi - February, 2026
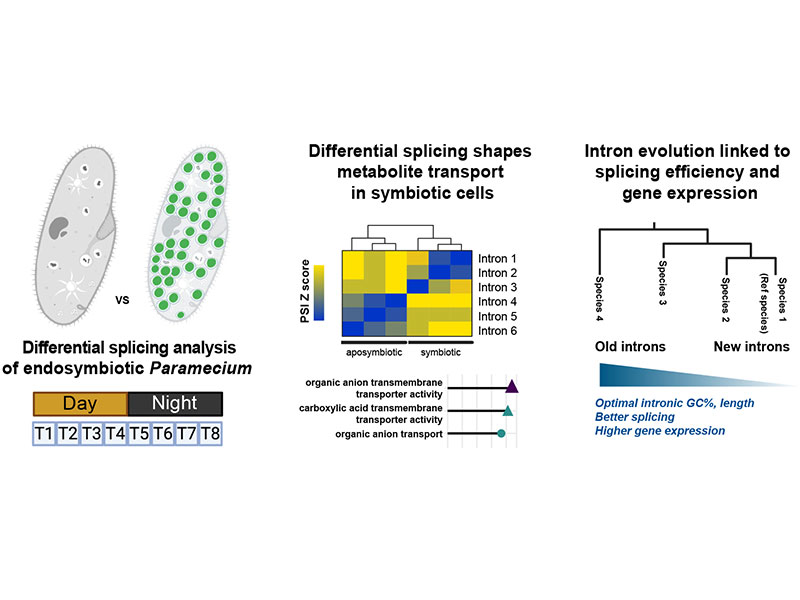
The integration of symbionts into host cells during endosymbiosis significantly alters gene expression and cell physiology. Though alternative splicing facilitates cellular adaptation through rapid modulation of gene expression and protein isoform diversity, its regulatory role during endosymbiosis remains poorly understood. Paramecium bursaria, which harbors hundreds of Chlorella variabilis algae within its cytoplasm, offers a powerful model to study splicing during endosymbiosis, especially given its exceptionally short introns (median ∼24 nt). Using time-course RNA sequencing of symbiotic and aposymbiotic cells, we found that splicing, especially of 5′ proximal introns, enhances gene expression. Moreover, we identified 883 genes with differentially spliced introns, particularly enriched in transmembrane transporters essential for establishing nutrient exchange between a host cell and algal symbionts. Splicing regulation correlated with expression changes in conserved spliceosome components, implicating that these factors act as splicing enhancers or repressors during symbiosis. By exploring intron orthology across ciliates, we found that conserved introns exhibited more efficient splicing, characterized by lower GC content and uniform length, suggesting that intron evolution favors features that optimize expression. Our study reveals how splicing contributes to host adaptation during endosymbiosis and highlights the evolutionary dynamics of short introns in eukaryotes.
How a Single-Celled Paramecium Rewrites Its Genes to Master Symbiotic Life
Distinguished Research Fellow Jun-Yi Leu and Assistant Research Fellow Chien-Ling Lin of the Institute of Molecular Biology led a research team integrating symbiotic evolution and RNA molecular biology to investigate how the ciliate Paramecium bursaria adapts to life with intracellular green algal symbionts through RNA splicing. RNA splicing is the process by which cells remove non-coding segments (introns) from transcribed RNA and join together the coding regions to produce functional messages. Notably, P. bursaria possesses exceptionally short introns, making it an ideal model for studying splicing and intron evolution. The study found that under symbiotic conditions, alternative splicing patterns of genes involved in nutrient transport are altered, indicating that splicing serves not only as a processing step but also as a regulatory mechanism that helps the host adapt to symbiotic living. Furthermore, these short introns exhibit specific sequence features that enhance splicing efficiency, suggesting that evolution can optimize gene architecture to improve organismal fitness in specialized environments.
The leading author of this study is Thi Ngan Giang Nguyen, a graduate of the Master’s Program in Genome and Systems Biology jointly organized by National Taiwan University and the Institute of Molecular Biology. This research was supported by the Academia Sinica Grand Challenge Program Seed Grant.
草履蟲如何「改寫」基因與綠藻共生
本院分子生物研究所呂俊毅特聘研究員與林倩伶助研究員的研究團隊結合共生演化與RNA分子生物學的研究,探討草履蟲 (Paramecium bursaria) 如何利用 RNA 剪接機制來適應與細胞內共生綠藻的生活。RNA 剪接是細胞把基因轉錄出的 RNA 中「不需要的片段」(intron)剪除、再把有用片段接起來的過程,而相對於其他物種,草履蟲的 intron 特別短,是研究剪接與演化的理想模型。研究發現,在共生狀態下,與養分運輸相關的基因剪接方式會發生變化,顯示剪接不只是基因加工步驟,更能調控基因功能,幫助細胞適應共生生活。同時,這些短 intron 具有特定的序列特徵,使剪接更有效率,說明演化會優化基因結構,以提升生物在特定環境中的生存能力。
本研究的第一作者是台大與分生所合辦的基因體與系統生物學程的碩士班畢業生Thi Ngan Giang Nguyen。研究經費由本院關鍵種子計畫支持。